Use Case 17 – Reusing data for validation of force fields

Quasi-elastic neutron scattering (QENS) is unique in the sense that it probes nuclei dynamics at the same time-scales that can be probed with molecular dynamics simulations. QENS data therefore offer an opportunity to validate force fields for molecular dynamics simulations and to optimize those force fields. This can be done simply by comparing the dynamic structure factor, calculated from molecular dynamics simulations with the corresponding one from real experiments. An example of that is given in the Figure below where experimental and simulated data are compared for two commonly used water models TIP3P and TIP4P-FB [1].
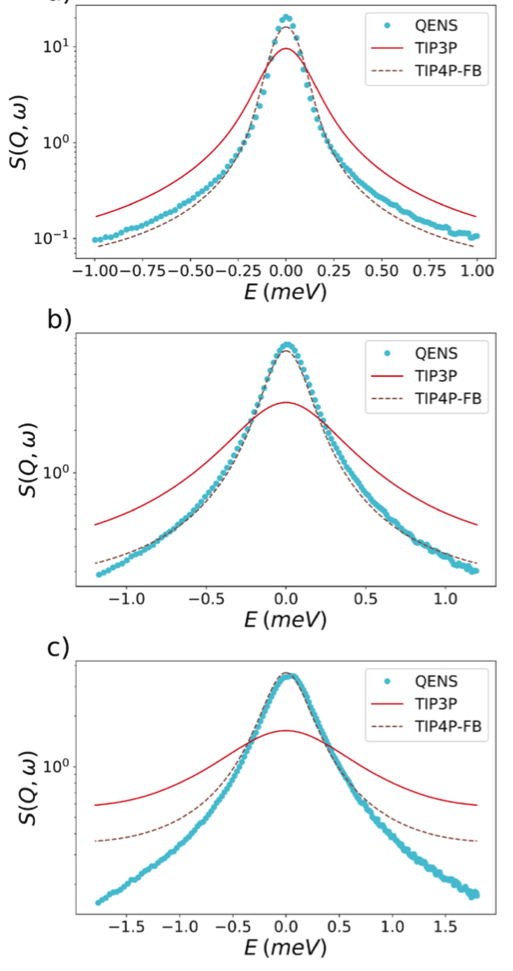
The experimental data shown in the above figure were acquired by a team [2] different from the one doing the molecular dynamics simulations and making the comparison in Ref. [1] and is, hence, also an example of data reuse.
In a second step, the force field can be optimized against the experimental data but this is not considered further in this use case.
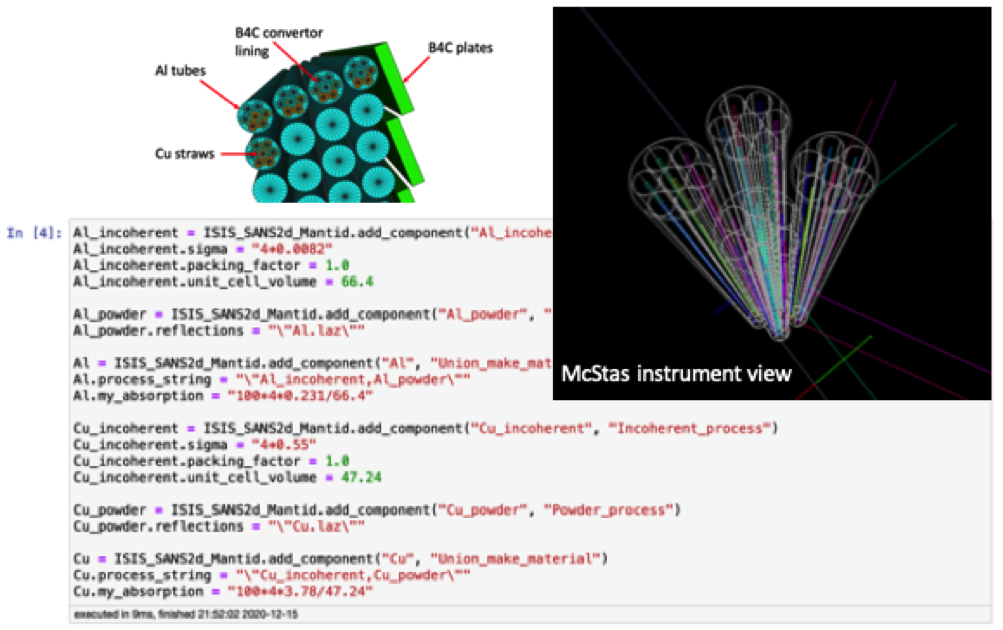
A desired outcome from PaNOSC /ExPaNDS would be the ability to search across all meta-data catalogues for specific types of samples (e.g. solvated amino acids) studied with a specific technique (e.g. QENS) or in a specific time- or energy range (e.g. meV) and retrieve the data incl. sufficient information (meta-data) so that the sample can be modeled in a molecular dynamics simulation. In this way, it would become possible to reproduce data for validating force fields for systems that a researcher would like to study with atomistic modelling and simulations and to explain experimental data through a detailed atomistic model. Ideally, data are auto-reduced and detailed information about the experimental setup (instrument and sample environment) is sufficiently detailed and presented so that experimental artefacts can be accounted for either through a resolution function or instrument / experiment simulations as illustrated below with a McStas simulation using McStasScript (PaNOSC WP5) of the ESS LoKI detector.

Main Contributors
- Prof. Jan Swenson, Chalmers University of Copenhagen – PI in the MDMC project funded by the Swedish Research Council and Tillväxtverket.
- Dr. Heloisa Bordallo, University of Copenhagen – Partner in the MDMC project
- Dr. Thomas H. Rod, ESS – Partner in the MDMC project
- Dr. Anders Markvardsen, ISIS – Lead developer in the MDMC project
Use Case Action Flow
- Inquiring: As a scientist, I inquire a service (web or desktop application, CLI or GUI) about specific data using keywords describing the composition and state of the systems of interest so that I retrieve an overview of accessible and matching data across multiple photon and neutron sources.
- Response: The system returns an overview of matching accessible data across multiple photon and neutron sources.
- Selection of data sets: I browse and select the data sets of interest for my project.
- Download reduced data sets: I download the associated reduced data ready for plotting and associated meta data, including appropriate resolution functions and instrument and sample environment geometries. Alternatively, I download the raw data to a system where I can process the data.
- Creating models: From the retrieved meta data I create models that mimic the samples. Meta data are presented so this step can be highly automated.
- Running simulations: I run the molecular dynamics simulations on a compute service using for instance the MDMC software.
- Conversion to scattering sections: I convert the trajectories from the molecular dynamics simulations to dynamical scattering cross sections folded with appropriate resolution functions and compare with experimental data.
- Simulation of experimental artifacts: As an additional step, as an alternative to using the resolution functions, I run instrument simulations with detailed description of the instrument(s), incl. detector geometry, and sample environment using the calculated cross sections for describing the sample. In practice, a digital twin of the experiment. For this step to be traceable, the process of going from instrument and sample environment description to instrument simulation should be semi-automated.
Description of needs
Step 1-4 require a service (data portal) that enables federated search across as many meta data catalogues as possible (PaNOSC WP3 & WP4) and auto-reduce the data (PaNOSC WP4).
Step 6 (and 7) requires a compute service with the appropriate software installed (or easy to install).
Step 8: Requires detailed models of instruments and sample environment and flexible instrument simulation software (e.g. McStasScript, PaNOSC WP5).
Impacts from the implementation
This would directly link the photon and neutron scattering science to that of materials and molecular modelling and simulations and:
- Potentially build up a new user community for the photon and neutron sources
- Expand the trust and usage of materials and molecular modelling and simulations through the opportunity to quantitatively validate the models.
- Make it possible to explain experimental data through a detailed atomistic description.
Generalisation of the use case
QENS, in relation to molecular dynamics simulations, is relevant for all neutron sources with QENS instruments. The concept is readily transferable to other techniques, e.g. diffraction across both photon and neutron sources using e.g. density functional theory for the modelling or Monte Carlo methods (see for instance RMC and EPSR), or time-dependent density functional theory in the case of spectroscopy at photon sources.
Resources
All software required is open source or will be made open source. The MDMC software will be made open source after it has been published.
Contact person
Thomas Holm Rod (European Spallation Source)
References
- [1] T.O. Farmer, A.J. Markvardsen, T.H. Rod, H.N. Bordallo, and J. Swenson. “Dynamical Accuracy of Water Models on Supercooling”, J. Phys. Chem. Lett. 11, 7469-7475 (2020). https://dx.doi.org/10.1021/acs.jpclett.0c02158
- [2] J. Qvist, H. Schober, and B. Halle. “Structural dynamics of supercooled water from quasielastic neutron scattering and molecular simulations. J. Chem. Phys. 134, 144508 (2011). https://doi.org/10.1063/1.3578472